Extending roxygen2
Maëlle Salmon (rOpenSci, cynkra)

Extending roxygen2
What I learned through use cases.
What is roxygen2?
Through roxygen2::roxygenize()/devtools::document():
Parsing of code decorators (tags) like
@detailsGeneration of manual pages and
NAMESPACE.
It is extensible.
It has been extended.
Example: devtag
devtag by Antoine Fabri (cynkra)
Document internal functions with roxygen2 tags and…
roxygen2’s
@noRd😭roxygen2’s
@keywords internal😕devtag’s
@devRd only for developers!.Rbuildignore🎉
Example: srr
srr by Mark Padgham (rOpenSci) to document compliance with statistical software standards.
Example: eDNAjoint by Abigail G. Keller
Example: roxytest
roxytest by Mikkel Meyer Andersen.
Inline tests with roxygen2 using testthat or tinytest.
“A big thanks goes to Mikkel Meyer Andersen for starting on the vignette and motivating me to make the extension process much more pleasant.” Hadley Wickham, roxygen2 7.0.0
Example: igraph
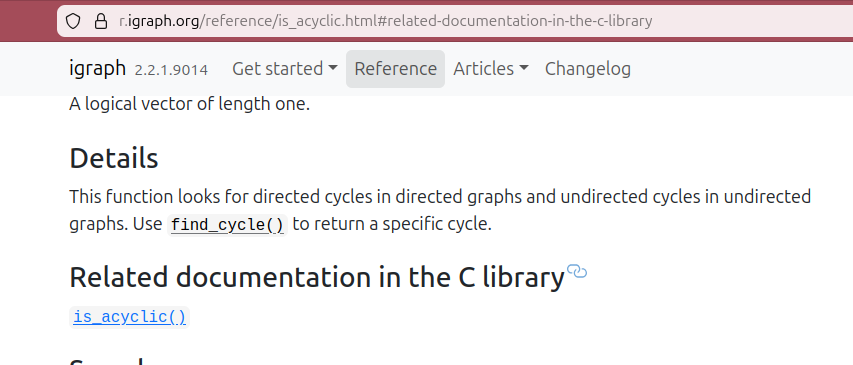
Example: vcr (WIP)
?ellmer::chat_openai without having any OpenAI account.
My use cases
✨ better docs for little effort ✨
I only want to create custom sections in Rd files. Easy, right?! 🥺
Extending roxygen2 (article)
There are two primary ways to extend roxygen2:
- Add a new tag that generates a new top-level section in .Rd files.
- Add a new roclet that does anything you like.
Person with an intense gaze explaining something using a corkboard covered by paper pieces and yarn connecting them
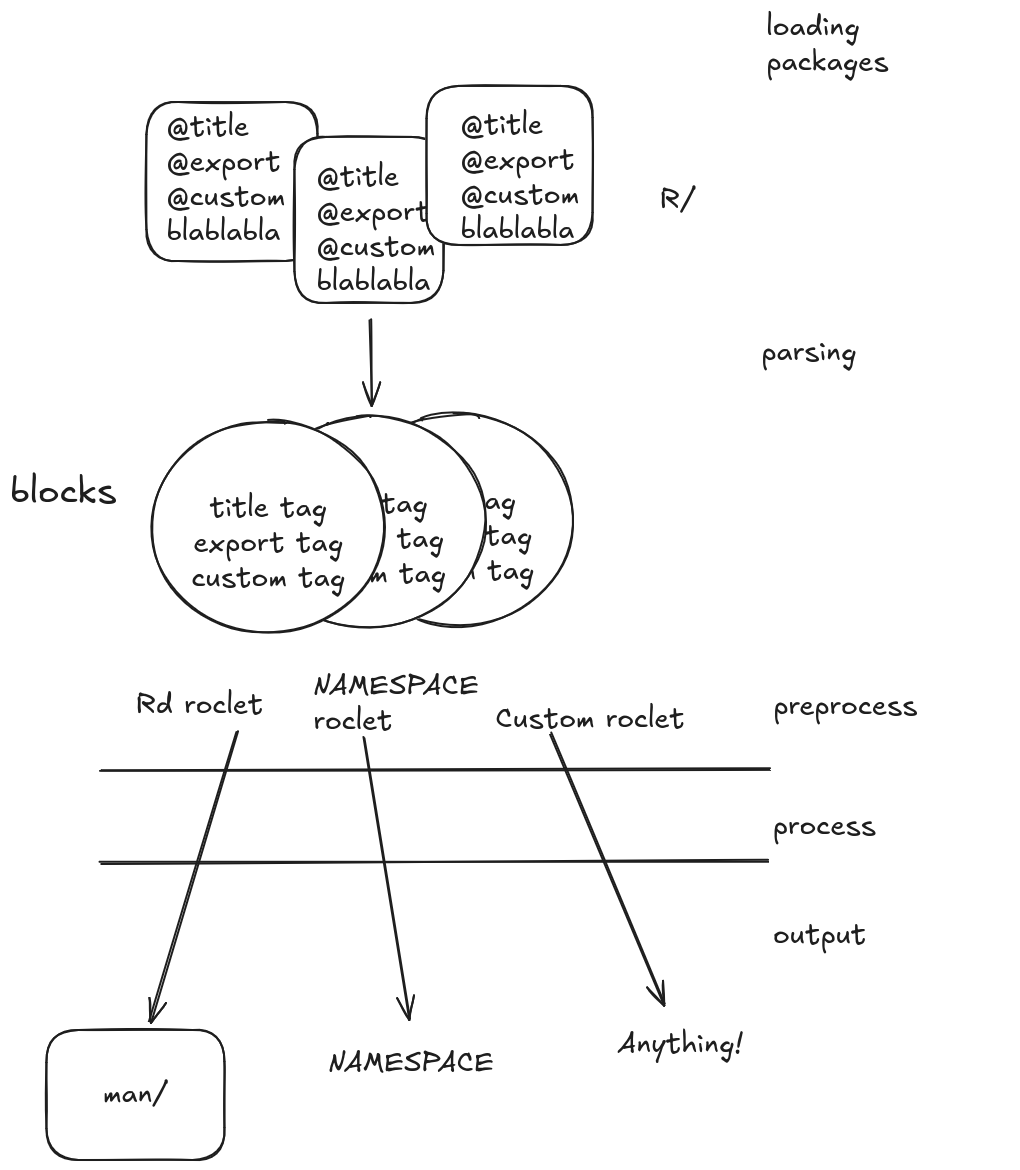
What roxygenize() does
Loads packages you tell it to load.
Parses all package R files, using available tags.
Finds the roclets you tell it to run. Rd roclet, NAMESPACE roclet, your roclet…
Runs the different methods of all those roclets. Preprocess, process, output.
parsing: If you define how to parse a tag roxygen2 will use it!! 🤯
roclet (S3 vessel for code!): roxygen2 will run your code!! 🤯
How to create a new tag: cdocs
the tag in use:
#' @cdocs igraph_is_acyclicthe Rd section rendered on the pkgdown website ✨
How to create a new tag for Rd files
roxy_tag_parse()method.roxygen2::tag_words()and friends. If absent: warning, tag ignored by parsing.
roxy_tag_rd()method.roxygen2::rd_section("cdocs", x$val)If absent: tag ignored by the Rd making.
- [If custom section]
format()method. Rd string creation.
How to share a new tag
Tag-creator package: Export the methods.
igraph.r2cdocs: Imports roxygen2 +
@exportggplot2 (
@aesthetics): Suggests roxygen2 +rlang::on_load()+vctrs::s3_register()
How to use a new tag
User package: Declare the use of the tag-creator package in DESCRIPTION.
packages
: packages to load that implement new tags.
https://roxygen2.r-lib.org/reference/load_options.html#possible-options
loadNamespace()
How to test a tag
Informally, obviously 😸
Beyond tags
Still focussed on Rd sections but…
A new roxygen2 extension for igraph
Less work please!
Get link to C docs without adding the @cdocs tag.
A new roxygen2 extension for igraph
For each manual page…
Get the aliases.
For each alias, get the call tree ({treesitter} FTW), extract names of C functions.
Create section with C links.
The challenge: An Rd section without tag!
roxygenlabs
https://github.com/gaborcsardi/roxygenlabs/
💡 Extend the Rd roclet
Use that roclet in lieu of the Rd roclet.
How to use/share a roclet
Roclet creator package: Export the roclet and its methods.
User package: Roxygen2 option to be stored in DESCRIPTION
roclets
: giving names of roclets to run.
About DESCRIPTION options
Roxygen field (to be retired?), maybe Config/Needs or Suggests field.
Documentation
Function like
use_devtag(), relying on {desc}
Summary: extending roxygen2 for Rd sections
Better docs with less manual work.
New tag. From a string to a templated section. Defined by methods.
packagesroxygen2 option.New roclet extending the Rd roclet. From information in the topic to a templated section. Defined by methods.
rocletsroxygen2 option.
Extending roxygen2 for anything
Anything a tiny bit fancy requires defining a roclet.
Could it be easier?

plumber2
https://github.com/posit-dev/plumber2/
api()instead ofroxygenize()/document()
Using/extending roxygen2
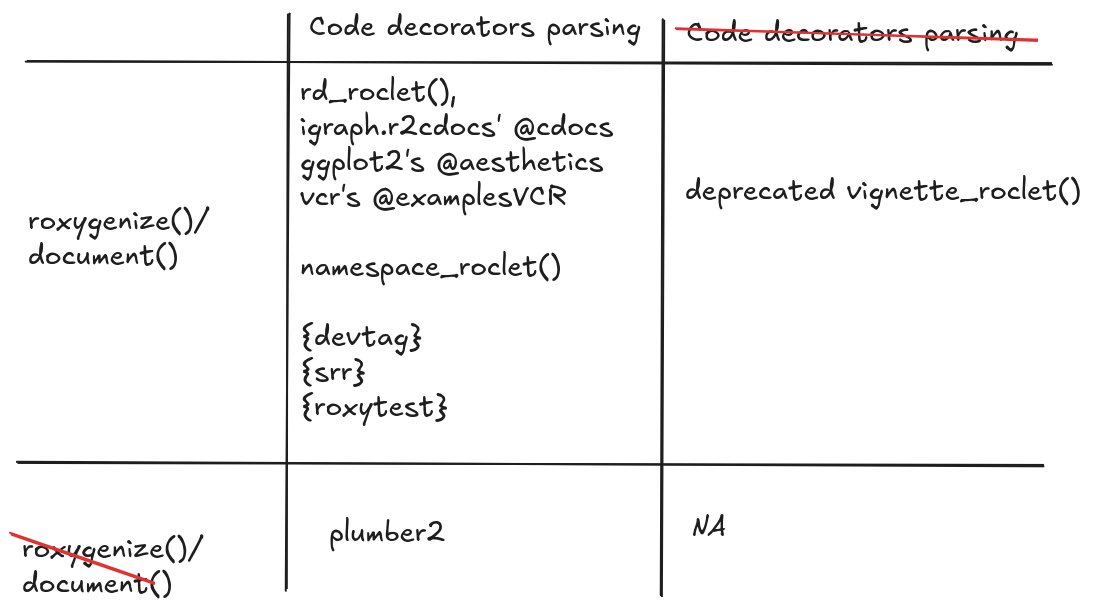
What to do?
to get good docs: read the roxygen2 docs website.
to repeat yourself less for Rd: create a custom tag.
to repeat yourself even less for Rd: extend the Rd roclet.
to do anything for packages: create a custom roclet.
to use code decorators outside of packages: use the parsing like plumber2.
roxygenize()
Conclusion
Extending roxygen2 is possible and useful!
Documentation still patchy or a bit hidden.
- please link to documentation explaining exactly what a roclet is, or omit the term from the documentation
- document how to load package with only extra Rd tags
- Consider adding some “advanced” manual pages to the reference index
- Document
subclassparameter ofroclet() - How to use custom options in DESCRIPTION for custom roxygen2 roclets?
Thank you!! 🙏
